The need for protein markers to distinguish particles from EVs
Accurate measurement of extracellular vesicles (EVs) in biofluids is essential for optimizing EV isolation workflow, EV biomarker discovery, and translational research. Label-free single-particle methods (e.g., flow cytometry, NTA, TRPS/MRPS) are the go-to methods for quantifying EVs. These methods detect particles within a defined size range but cannot distinguish vesicles from other nano-sized components commonly present in biological fluids. In complex biofluids such as plasma or serum, lipoproteins and protein aggregates often overlap with EVs in both size and density, complicating accurate measurement (1-5). MISEV2023 now formally distinguishes “particle concentration” from “EV concentration” and recommends orthogonal protein-based methods to close this gap (1). Specificity of single particle techniques can be improved through orthogonal detection of EV-associated proteins (e.g. CD9, CD81, CD63) through fluorescent labeling, which are commonly enriched on the EV membrane. Fluorescent labeling of EVs for single-particle analysis requires optimization, extensive controls, and is limited by sensitivity.
Western blot for EV markers: Orthogonal confirmation with broad limitations
Western blot, an inherently multi-step and laborious method, has traditionally been used to confirm canonical EV markers such as CD9, CD63 and CD81. By separating proteins by molecular weight, it provides an additional layer of molecular identity verification. However, because it analyzes denatured protein, Western blot cannot distinguish EV-associated markers from non-vesicular forms, a critical limitation when free protein contamination is high. Performance also declines in complex samples such as plasma, resulting in smeared bands that are difficult to resolve. Furthermore, Western blot has limited sensitivity and cannot detect low levels of proteins. These limitations highlight the need for a more robust, sensitive, and scalable approach.
ELISA-based immunoassays can address workflow and sensitivity limitations of Western blot. With same-day readouts and plate-based throughput, ELISA enables reproducible, efficient EV analysis across complex biofluids. Traditionally, ELISA assays are designed to detect soluble proteins, which do not provide EV-specificity. In the table below, we compare Western blot and ELISA across key parameters relevant to EV protein analysis (1, 6-8).
| Parameter | ELISA | Western Blot |
| Detection principle | Antibody capture + enzymatic signal amplification | Electrophoresis + antibody detection |
| Detection context | Native proteins | Denatured proteins after SDS-PAGE |
| Analytical sensitivity | High –moderate (depending on readout method) | Low |
| Dynamic range | 2-4 logs (depending on readout method) | Limited |
| Specificity | Antibody pair–dependent | Antibody binding with molecular weight confirmation |
| Performance in complex biofluids | Compatible with plasma, CSF, urine, saliva, conditioned media | Reduced in plasma due to high protein background |
| Time to result | Same-day (~hours) | ~1.5–2 days |
| Workflow complexity | Simple; 1-3 incubation steps + wash | Multi-step (electrophoresis, transfer, antibody incubations) |
| Throughput/scalability | High; plate-based, automation compatible | Low; manual lane-based workflow |
Combining Western Blot and ELISA for complementary EV analysis
In practice, ELISA and Western blot are most effective when used together. Ter-Ovanesyan et al. showed this clearly when optimizing the immuno-isolation of neuron-specific EVs in plasma. ELISA measured tetraspanin depletion in the supernatant, whereas Western blot produced only a smear. Western blot confirmed target enrichment in the pulldown, which cannot be quantified by ELISA because measuring bead-bound antigen requires disrupting the antigen–antibody interaction to elute the target (9). Taken together, Western blot confirms EV marker presence, and ELISA confirms its depletion from the supernatant, allowing for EV subtype pull-down optimization.
Atlas EV ELISA: Multi-target intact EV ELISA for EV quantification and characterization
To overcome the challenges of traditional ELISA for EV quantification, Everest Biolabs developed the Atlas EV ELISA, an intact EV assay that simultaneously targets the three most abundant tetraspanins: CD81, CD63, and CD9. This plate-based format quantifies EV-associated proteins and evaluates sample purity using only 50 μL of biofluid or SEC fraction with just 15 minutes of hands-on preparation. The Atlas Human EV ELISA enables comparison of EV yields and standardization of inputs for downstream studies. For example, Atlas absorbance ELISAs are particularly powerful for quantifying EV yield and purity across SEC fractions, thereby fine-tuning fraction collection and optimizing workflows for downstream applications. More on this.
The Atlas Lumi Human EV ELISA, which uses chemiluminescence readout, provides higher-sensitivity and wider-dynamic range necessary for low-abundance samples such as CSF or dilute culture media. For researchers targeting markers beyond the tetraspanins, the Atlas Lumi Homebrew offers a customizable format to pair the assay with proprietary antibodies.
Schematics of the workflow for Western Blot, ELISA and Atlas EV ELISA for protein detection
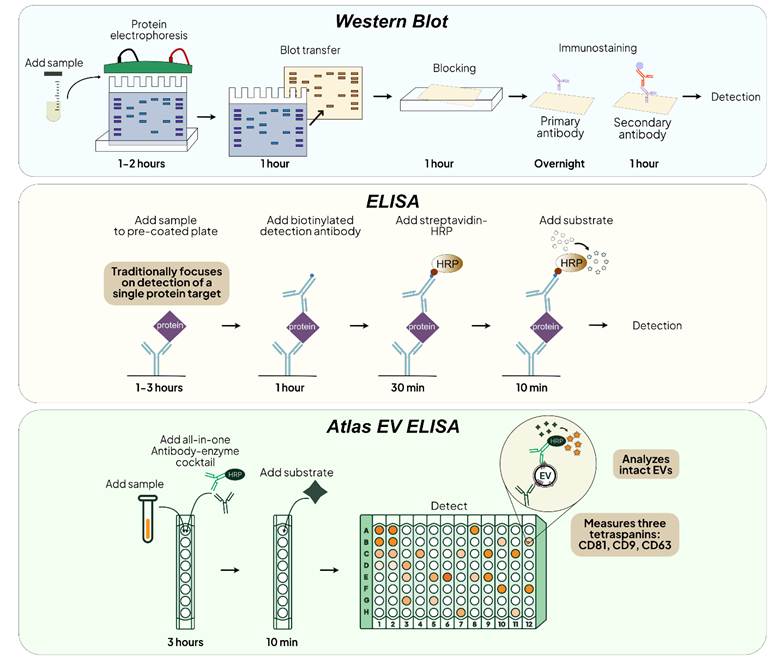
Summary
Label-free single-particle detection techniques provide the total particle concentration and are not EV-specific. Western blot is widely used to confirm the presence of canonical EV markers such as CD9, CD63, and CD81 through orthogonal affinity- and molecular weight-based detection, but it provides limited quantitative or surface-specific insight into EV populations. In contrast, Atlas EV ELISA measurement of surface markers on intact vesicles enables scalable, quantitative, and biologically interpretable assessment of EV abundance and variation across biofluids and clinical states. Together, these distinctions position Western blot primarily as a confirmatory identity tool, while EV ELISA serves as a more suitable platform for sensitive, reproducible, and cohort-scale EV biomarker measurement.
References
- Welsh, J.A., et al. (2024). Minimal information for studies of extracellular vesicles (MISEV2023). J Extracell Vesicles, 13(2): e12404.
- Welsh, J.A., et al. (2017). Extracellular vesicle flow cytometry analysis and standardization. Frontiers in Cell and Developmental Biology, 5: 78.
- Welsh, J.A., et al. (2022). MIFlowCyt-EV: a framework for standardized reporting of extracellular vesicle flow cytometry experiments. Journal of Extracellular Vesicles, 11(2): e12182.
- Mørk, M., et al. (2017). Prospects and limitations of nanoparticle tracking analysis (NTA) in extracellular vesicle research. Journal of Extracellular Vesicles, 6(1): 1305676.
- Davidson, S.M., et al. (2023). Extracellular vesicles and cardiovascular disease: biology and translational opportunities. Cardiovascular Research, 119(5): 1132–1148.
- Kowal, E.J.K., et al. (2017). Extracellular Vesicle Isolation and Analysis by Western Blotting. Methods in Molecular Biology, 1660: 143–152.
- Logozzi, M., et al. (2020). Immunocapture-based ELISA to characterize and quantify exosomes in both cell culture supernatants and body fluids. Methods in Enzymology, 645: 155–180.
- Hartjes, T.A., et al. (2019). Extracellular Vesicle Quantification and Characterization: Common Methods and Emerging Approaches. Bioengineering (Basel), 6(1): 7. PMID: 30654439.
- Ter-Ovanesyan, D., et al. (2024). Identification of markers for the isolation of neuron-specific extracellular vesicles. bioRxiv.